 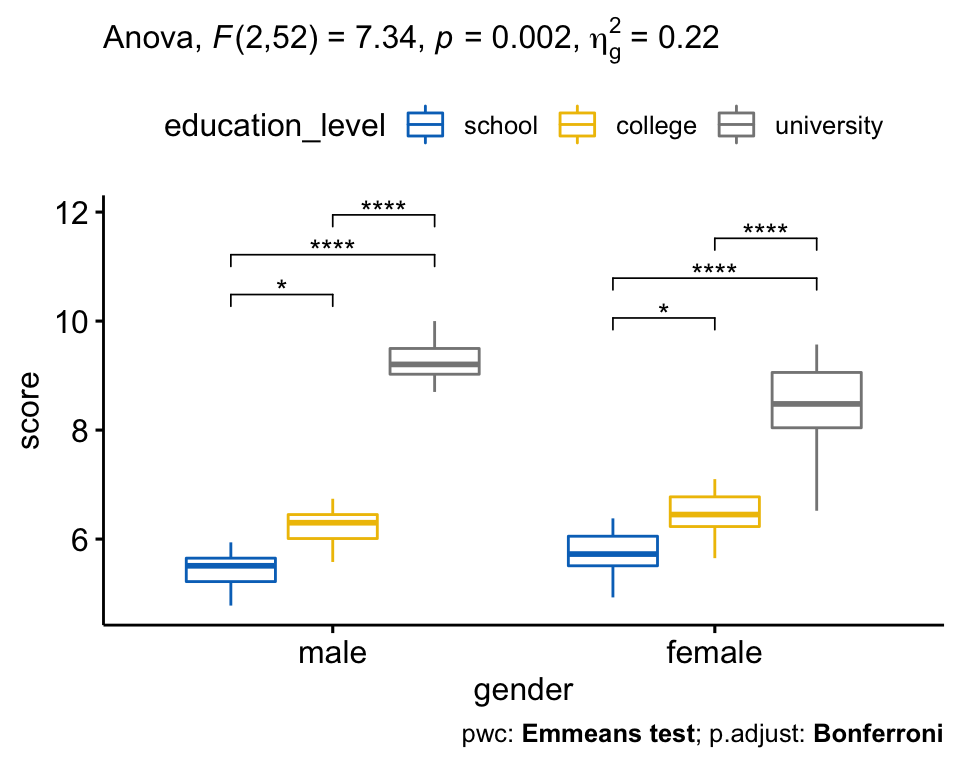 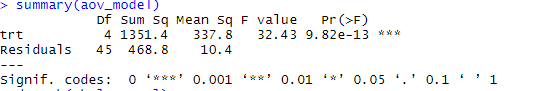 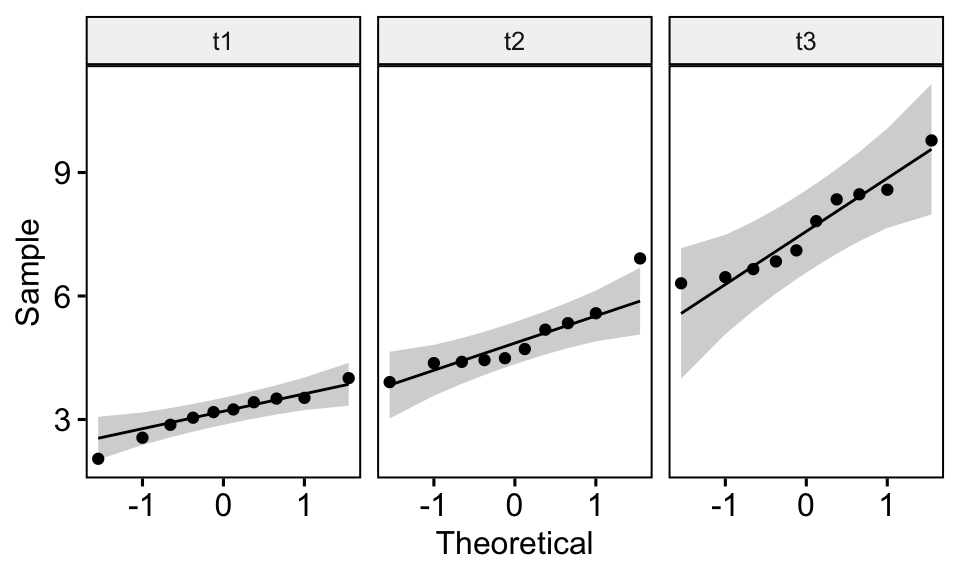
TukeyHSD(weeds.aov) # Tukey multiple comparisons of means# 95% family-wise confidence level# Fit: aov(formula = flowers ~ species, data = weeds)# $species# diff lwr upr p adj# Olearia-Coprosma 12. But without conducting an extra test, we cannot be certain which species are statistically significant from each other when it comes to their effect on flower abundance codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1This example constructs an ANOVA with two factors, but does not include the interaction term.Īll of our analyses so far have showed us that species has an influence on flower abundance. To conduct an two-factor ANOVA is pretty straightforward. ANOVA is one of the most basic yet powerful statistical models you have at your disopsal. This chapter describes methods for checking the homogeneity of variances test in R across two or more groups. The normal way to do so is to use the anova() command.Īnova(weeds.aov) # run an anova on the object# Analysis of Variance Table# Response: flowers# Df Sum Sq Mean Sq F value Pr(>F) # species 2 2368.6 1184. ANOVA (or AOV) is short for ANalysis Of VAriance. Once we know our data is normal and we have our aov() object, we can use one of two commands on this object to generate our statistical result. Any violations of these assumptions can cause the test to produce a false-positive, as an analysis of variance is sensitive to violations in the assumptions of normality and homogeneity of variance (also called homoscedasticity). This is due to the algorithms at work assuming your data fits specific distributions, has equal variances and your replicates are independant of one another (not spatially autocorrelated). Content:Īnalyse the relationship between mutliple categorical predictor variables on a continuous response variableĪll statistical tests need to make various assumptions about your data when conducting the test. These will be specified within a “instruction” block before the requisite code chunk. In these tutorials, we will be using a variety of datasets to test each analysis. Procedure is quite simple for One-Way ANOVA: bartlett.test (x g) where x is numeric, and g is a factor var.test (x g) But, for 2x2 tables, i.e. The F ratio is then compared to a critical value from a table of values (based of degrees of freedom) to determine the level of significance, or P value. I've been using var.test and bartlett.test to check basic ANOVA assumptions, among others, homoscedascity (homogeniety, equality of variances). The variance for each point is squared and added together to generate the sum of squares, which is then used to generate the F ratio. The ANOVA tests the difference between the factors variance (distance from the grand mean) compared to the error variance. An ANOVA is used to test the effect of 1 or more categorical explanatory variables (X) on a continuous response variable (Y). Repeated measures, non-parametric, multivariate analysis of variance as far as I know, such a method is not currently available in R. To begin our foray into statistics in R, we will start with the most basic and useful analysis, Analysis of Variance (ANOVA). Built with the "Learn" Theme using Hugo and Blogdown 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
